
Improving ultrasound video classification: an evaluation of novel deep learning methods in echocardiography
Introduction
Echocardiography is the commonest use of medical ultrasound, and efforts are being made to streamline the time-consuming process of reporting and automate the analysis of studies to allow sonographers to scan more patients. One major barrier, however, is that each study comprises upwards of 50 video loops providing depictions of slices of the heart in a variety of anatomical planes and orientations, also known as ‘views’. Before each video loop can be analysed, the view it represents must be correctly identified. One possible approach to this task is using neural networks, which have been employed successfully in several other fields of medical image classification (1-3).
However, the accuracy of echocardiogram view classification using neural networks has been variable (4,5) with studies typically classifying individual video frames in isolation, missing the opportunity to use temporal information of how features such as heart valves or ventricular walls move during the cardiac cycle.
In this paper we first assess the relative performance of several classical convolutional neural network (CNN) architectures. We then introduce some new architectures, inspired by current work in the field of human action recognition, which can process both the spatial and temporal information contained in video (Figure 1). The four groups of architectures we asses are as follows:
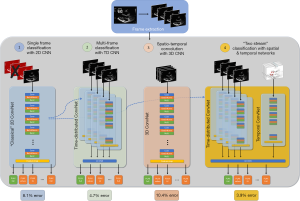
- Classical CNN. We assessed 5 different CNN architectures each of which has held the title of being the state-of-the-art in network design for image recognition. As with previous studies, accuracy is calculated using the modal prediction across multiple video frames (4).
- Classical CNN encapsulated within a time-distributed layer. In this design, an entire video is passed through a classical CNN, frame by frame, and CNN’s output from each frame is collated. Then each output is fed, sequentially, into a second neural network.
- Three-dimensional (3D) CNN. We assessed the ability of a 3D CNN which comprises filters which not only scan (‘convolve’) through the two dimensions of each frame of a video, but also across frames in the third dimension.
- “Two stream” CNNs. This network takes in two streams of data: a spatial stream, and a temporal stream. The spatial stream processes sequential video frames and comprises either a time-distributed classical CNN network (see 2) or 3D CNN (see 3). A second ‘temporal’ network is also trained to which receives optical flow data, i.e., data describing the movement of objects between frames. The final view decision is based on integration of both temporal and spatial signals.
Finally, we compare the error rate of the best performing network with the disagreement rate of two echocardiography experts, which may represent the upper limits of what is achievable using retrospective human-labelled data.
Methods
Data extraction
A random sample of echocardiogram studies and their associated video loops were extracted from Imperial College Healthcare NHS Trust’s echocardiogram database in DICOM format. Ethical approval was gained from the Health Regulatory Agency (Integrated Research Application System identifier 243023). Only studies with full patient demographic data and without intravenous contrast administration were included. All videos comprising at least 40 frames were analysed. Automated anonymisation was performed by identifying the ultrasound machine model from the DICOM file meta-data and blanking the pixel range in which that model displays patient-identifiable information. Individual studies were randomised to the training or testing sets at a ratio of 3:1.
An expert human classified each video from the training and test sets into one of 14 categories which are outlined in Figure 2. Videos thought to show no identifiable echocardiographic features, or which depicted more than one view, were excluded. These classifications (labels) were used to train and assess the performance of the network. A second human classified each video from the test set, unaware of the other human’s classifications, so that inter-expert agreement could be assessed.
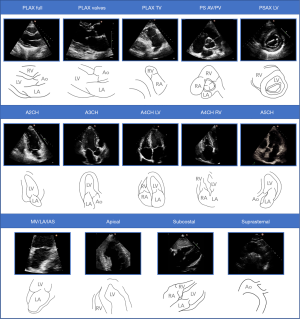
The first 40 frames of each video were then extracted and resized to a resolution of either 299 by 299 or 224 by 224 pixels, depending on the requirements of the network being trained. Optical flow frames were extracted from videos using denseflow (6) at a resolution of 224 by 224 pixels. Images were normalised at training time to yield pixel values between 0 and 1. The effects of hyperparameter tuning and training progress were assessed during training using 20% of the training set as a ‘validation’ subset on which the network was not trained.
Detailed information regarding the neural network architectures, training and statistical analysis are available in Supplementary file.
Results
Dataset
Three hundred and seventy-four echocardiographic studies met the inclusion criteria for extraction. They were randomly split into the training (75%) and testing (25%) datasets. Together, they contained 9,098 echocardiographic videos. Of these, 8,732 (96.0%) videos could be classified as one of the 14 views by the first expert. The remaining 366 videos were not classifiable as a single view, either because the view changed during the video loop, or because the images were completely unrecognisable. The study and video characteristics are shown in Table 1.
Table 1
| Characteristics | Training set | Testing set |
|---|---|---|
| Study characteristics | n=282 | n=92 |
| Sex (male) | 144 (51.1) | 46 (50.0) |
| Age | 61.7 (18.6) | 61.8 (18.5) |
| Manufacturer | ||
| Philips | 224 (79.4) | 72 (78.3) |
| GE | 58 (20.6) | 20 (21.7) |
| Video characteristics | n=6,592 | n=2,140 |
| Colour flow Doppler | 718 (10.9) | 234 (10.9) |
| Classes | ||
| Parasternal long axis | 713 (10.8) | 260 (12.2) |
| Parasternal long axis valves | 426 (6.5) | 123 (5.7) |
| Parasternal long axis RV inflow | 207 (3.1) | 63 (2.9) |
| Parasternal short axis LV | 1,045 (15.9) | 343 (16.0) |
| Parasternal aortic & pulmonary | 874 (13.3) | 291 (13.6) |
| Apical 2 chamber | 534 (8.1) | 152 (7.1) |
| Apical 3 chamber | 503 (7.6) | 145 (6.8) |
| Apical 4 chamber LV | 829 (12.6) | 289 (13.5) |
| Apical 4 chamber RV | 222 (3.4) | 79 (3.7) |
| Apical 5 chamber | 295 (4.5) | 84 (3.9) |
| Apical LA/MV focused | 199 (3.0) | 58 (2.7) |
| Apex | 25 (0.4) | 9 (0.4) |
| Subcostal | 570 (8.6) | 193 (9.0) |
| Suprasternal | 150 (2.3) | 51 (2.3) |
| Unclassifiable* | 273 (NA) | 93 (NA) |
Categorical data is shown as numbers (%); continuous data are shown as mean (standard deviation). *, 273 and 93 videos from the training and testing sets, respectively, were unable to be classified, and do not contribute towards the class percentages; this is because they showed two or more views during the full video loop, or neither of the human operators were able to identify sufficient landmarks to classify the video. LV, left ventricle; RV, right ventricle.
Performance of the different architectures
The best-performing classical 2D CNN design (Table 2) was Xception, with an error rate of 8.1% (Cohen’s Kappa 0.910). The two distinctions that were most error-prone were (I) 20 cases of confusing A4CH with A5CH and (II) 19 cases of confusing PLAX inflow versus parasternal aortic and pulmonary valves.
Table 2
| Architecture | Trainable parameters (million) | % Error | Cohen’s kappa |
|---|---|---|---|
| Classical CNNs | |||
| VVGNet 16 | 14.7 | 86.4 | 0 |
| Inception V3 | 21.9 | 11.9 | 0.868 |
| Resnet | 23.6 | 11.0 | 0.878 |
| DenseNet 121 | 7.0 | 10.0 | 0.889 |
| Xception | 20.9 | 8.1 | 0.910 |
*, these figures reflect the accuracy of the temporal network alone, which goes on to form one stream of each of the two-stream networks assessed. CNN, convolutional neural network; 3D, three-dimensional; TD, time-distributed.
The time-distributed CNN performed better (P<0.0001) with an error rate of 4.7% (Cohen’s Kappa 0.947).
The two 3D CNNs examined gave disparate results. The modified C3D network performed poorly (Table 2) and failed to provide any meaningful classification. However, the I3D network yielded an accuracy superior to that of the 2D CNN on which it was based (Inception 3D; 10.6% error versus 11.9% error). However, this network proved inferior to time-distributed CNNs (P<0.0001).
The two-stream networks demonstrated the highest accuracies, with the network based on the time-distributed CNN having an error rate of only 3.9% (Cohen’s Kappa 0.957). This performance was superior to that of both the classical 2D CNNs and 3D CNNs (P<0.0001), though superiority versus the time-distributed CNN did not reach statistical significance (P=0.053). For this network, the two distinctions that remained most error-prone were (I) 10 cases of confusing A4CH with A5CH and (II) 8 cases of confusing A2CH with A3CH.
Confusion matrices for each class of network are shown in Figure 3A,B,C,D. The changes in classifications associated with using the best performing model, rather than a classical 2D CNN are shown in Figure 3E.
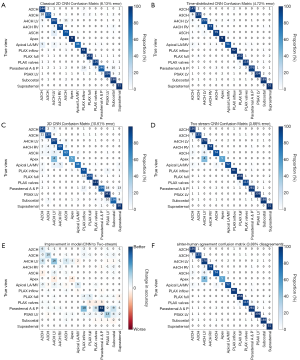
Inter-expert agreement
The results in Figure 3 show disagreement (or error) rates of only a few percent in the best performing networks, and these errors appear predominantly clustered between certain pairs of views which represent anatomically adjacent imaging planes. It was conceivable, therefore, that some of the residual apparent error was due to an inherent difficulty of deciding between views that are similar in appearance and are in spatial continuity.
To investigate this, the test set classifications of the second expert cardiologists were compared to those of the first expert.
Of the 2,140 test set videos, 74 (3.5%) were classified differently by the second expert (Figure 3F). The two distinctions that caused the most disagreement were (I) 13 cases of A5CH versus A4CH and (II) 10 cases of A3CH versus A2CH.
When allowing either expert classification as correct, the error rate decreased further, to 2.6%.
Finally, addressing only the 2,064 test videos classified identically by the two cardiologists, the error rate of the two-stream network was only 2.2%, corresponding with 45 videos “misclassified” by the network.
For reference, the two-stream network’s error rate when judged solely against the second human’s classification was 4.3%.
Discussion
This study shows that the application of new CNN architectures can reduce the error rate of echocardiographic video classification by more than two-fold. It further suggests the remaining error rate may contain a substantial element of “judgement calls”, where experts are uncertain and when forced to commit to a class independently, pick a different class from other experts.
Temporal neural networks achieve half the error rate of classical 2D CNNs
We had success with two approaches of integrating temporal information: time-distributed networks and two-stream networks. The two-stream network had an error rate less than half that of the best classical 2D CNN.
Much of this benefit appears to be through improved discrimination between certain pairs of views that classical CNNs find challenging. For example, 19 videos were misclassified by the 2D CNN, but only 2 by the two-stream network, when deciding between the ‘PLAX inflow’ and ‘parasternal aortic and pulmonary valves’ views. On a single frame (Figure 4A,B) these videos are easily confused, even by a human expert. One clue to their correct classification is the direction of motion of the leaflets when the valve opens: upwards versus downwards. In most static images it is almost impossible to distinguish, and the true identity only emerges when a temporal sequence of images is examined. The resulting error rate for this ‘parasternal aortic and pulmonary valves’ view fell from 17.5% for the classical 2D CNN to just 3.4% for the two-stream network.
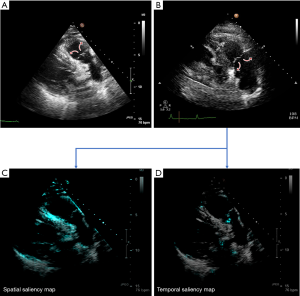
Figure 4C,D show saliency maps derived from the two-stream network for this parasternal pulmonary valve example. Saliency maps highlight the features of the videos which are contributing most towards the neural network’s decision. Figure 4C appears to indicate the spatial arm of the network’s decision is largely influenced by the anatomical borders of the major structures such as the pulmonary artery and left ventricle (highlighted in cyan). The temporal (optical flow) arm’s saliency map (Figure 4D), in contrast, shows intense focal activation over the pulmonary valve leaflets. These visualisations may support the theory that the two-stream network’s ability to discriminate between such classes may in part be due to their ability to track the movement of structures such as valves throughout the cardiac cycle.
Temporal neural networks show both a magnitude and distribution of errors comparable to humans
The time-distributed 2D CNN and two-stream network, which process each 2D image in a video sequentially, avoid the enormous resource consumption of 3D CNNs, which may be why they achieve such high accuracies.
Using one human expert as the reference, the best-performing neural network agreed in 96.1% of cases (3.9% error), while the second expert agreed in 96.4% (3.6% error). If the dataset is restricted to only those images where the two humans agreed, the network was 97.8% accurate, with only 45 (2.1%) video loops classified differently by the network.
Interestingly, the two views the network found most difficult to correctly categorise (Figure 3D) were the also the two views on which the two experts disagreed most often (Figure 3F): A4CH versus A5CH, and A2CH versus A3CH. The A4CH view is in an anatomical continuity with the A5CH view. The difference is whether the scanning plane has been tilted to bring the aortic valve into view, which would make it A5CH. When the valve is only partially in view, or only in view during part of the cardiac cycle, the decision becomes a judgement call and there is room for disagreement. Similarly, the A3CH view differs from the A2CH view only in a rotation of the probe anticlockwise, again to bring the aortic valve into view.
Study limitations
Interpreting the results of a neural network study alongside previous studies can prove difficult. There have been two previously published papers assessing the role of classical 2D CNNs for view classification, and they have published very different results, with accuracies of 84% (5) and 97.8% (4), respectively. There are several possible explanations for the wide range of reported accuracies.
First, the more numerous the view categories, the more difficult the task of the neural network, since if a group of videos are considered a single view in one study but multiple views in another, those multiple views are likely to be relatively similar in appearance. Moreover, one of the studies grouped all images possessing colour as a single category, regardless of the anatomical plane (5). Since identifying the presence of colour is simple, this increases the reported accuracy of any network.
Second, it is possible that previous studies included videos that were more easily differentiated as classical planes, which could explain complete lack of confusion in one study (4) between A2CH and A3CH, which are anatomically continuous.
Third, studies sometimes show networks performing better than humans, but have trained and tested the network on reduced-resolution images such as 80-by-60 pixels and have accordingly tested the humans on such low-resolution images, which they will not have had experience of distinguishing (4).
For these reasons, in this paper we have re-implemented 5 different classic 2D CNN architectures, and it is against these which the novel architectures have been assessed. Furthermore, the human experts were provided with full-resolution ultrasound videos, even though the networks were using reduced-resolution data.
Conclusions
In this study of over 8,000 echocardiographic videos, we have shown that switching to advanced neural network architectures can halve the error rate for view classification, reaching the concordance achieved by second opinions from blinded human experts. Moreover, the types of misclassification these advanced networks now make are very similar to the sources of differences of opinion between human experts.
Supplementary
Neural network details
For the classical 2D CNNs, we investigated the performance of 5 different neural network architectures: DenseNet, Inception V3, ResNet, VGGNet and Xception. Each network was initialised using weights derived from training on ImageNet, a large image database used for object recognition. The final convolutional layer of each network was fed into a global average pooling layer before the final output layer. During testing, 5 linearly spaced frames were chosen from each video and the modal answer was used when calculating accuracy. The best performing architecture (Xception) from this stage was used as the spatial convolutional base for the time-distributed and two stream networks.
The time-distributed 2D CNN comprised the trained Xception network which was fed the first 20 frames of each echocardiogram. The feature maps (from the global average pooling layer) for each frame were fed into three 1-dimensional depthwise-separable convolutional layers comprising 256 kernels of filter size 3, separated by 1-dimensional max pooling layers of pooling size 2. Finally, a 1-dimensional global average pooling layer interfaced with the final output layer. This architecture is similar to the “long-term recurrent convolutional networks” pioneered by Donahue et al. (7) but our design differs in several important respects. First, we used a series of depthwise-separable convolutional layers instead of long-short term memory layers. Second, we used a 1-dimensional global average pooling layer with dropout before the final output layer. We found that these two design decisions drastically reduced the number of parameters in the model and aim to minimise over-fitting whilst not impacting on accuracy.
Two 3D CNN architectures were assessed. The first 3D CNN was a modified form of the C3D architecture pioneered by Tran et al. (8) Given this architecture would require over 1 billion trainable parameters when passed images of the dimensions used in our study, we swapped the flattening layer for a 3D global average pooling layer which has since become the modern practice (9) and which also drastically reduces the number of parameters. The second 3D CNN was created using the Inception 3D architecture as published by Carreira et al. (10) The network was pretrained on ImageNet and Kinetics using weights released by Deepmind (11). Model weights were frozen for the first epoch before unfreezing weights progressively frontwards over 5 epochs to minimise catastrophic forgetting of learned weights during early gradient updates. Both networks received the first 20 frames of each echocardiogram.
Finally, the two-stream networks comprised two distinct convolutional ‘streams’ (one ‘spatial’ stream, one ‘temporal’ stream), which process a video’s spatial and temporal features separately before the data are integrated and a final decision of the view is made. The spatial stream comprised either a time-distributed 2D CNN or a 3D CNN, and was fed the first 20 frames of each echocardiogram. The temporal stream comprised a separate classical CNN architecture which was trained to identify videos using only optical flow data, comprising greyscale heatmaps highlighting the movement of structures between two sequential frames in a video. A single data sample fed into the temporal network was made up of data describing the inter-frame optical flow from a series of 10 frames, with a separate frame for movements in the vertical and horizontal planes, resulting in a 20-channel input image of 224 by 224 pixels for a 10 frame ‘chunk’. The temporal network was encapsulated in a time-distributed layer and followed by a 1-dimensional global average pooling layer, allowing 4 sets of 10 frame chunks within a video to be processed. Finally, the spatial and temporal global average pooling layers were concatenated before the final output layer. This design is inspired by the two-stream networks pioneered by Feichtenhofer et al. (12), but differs in several key respects, most notably that our temporal stream comprises an untrained Inception V3 (13) model. Unlike Feichtenhofer’s implementation, this network contains no fully connected layers and uses global average pooling of feature maps which results in significantly fewer trainable parameters and we found led to much faster convergence with improved accuracy.
Each network was trained until the validation loss plateaued. Models were saved after each epoch, and the model with the highest validation accuracy was used for the final assessment on the test set. The final output layer of every network comprised 14 densely-connected neurons (one for each view). Loss was calculated using the categorical cross entropy loss function and weights were updated using the Adam optimizer. The batch size for all networks was 20.
Saliency maps were used to further investigate the relative focus of the two components of the two-stream networks (spatial features and temporal features). To allow pictorial visualisation of this, despite the networks receiving input in video format, the saliency patterns across each spatial frame were averaged and then normalised, before being super-imposed upon a single frame from the video.
Programming was performed with the Python programming language, with the Tensorflow (14) and Keras (15) machine learning frameworks and the Keras-vis package (16).
Statistical analysis
The primary statistical endpoint was accuracy, defined as the proportion of videos correctly classified according to their view. Confidence intervals for accuracy were calculated using the “exact” binomial method. Significance testing between models was by McNemar’s test with P=0.05 as the threshold for statistical significance, with an exact test used for contingency tables including any counts below 25. Cohen’s kappa was calculated for each model to account for imbalanced class sizes. Statistical analysis was performed using the R programming language.
Supplementary
Acknowledgments
We thank Quasim Ahmed for infrastructure support.
Funding: JPH is funded by the Wellcome Trust (212183/Z/18/Z). Dr. Rueckert has received research grants from and is a consultant for Heartflow. This study was supported by the NIHR Imperial Biomedical Research Centre (BRC). The views expressed are those of the author(s) and not necessarily those of the NIHR or the Department of Health and Social Care.
Footnote
Conflicts of Interest: All authors have completed the ICMJE uniform disclosure form (available at http://dx.doi.org/10.21037/jmai.2019.10.03). The authors have no conflicts of interest to declare.
Ethical Statement: The authors are accountable for all aspects of the work in ensuring that questions related to the accuracy or integrity of any part of the work are appropriately investigated and resolved.
Open Access Statement: This is an Open Access article distributed in accordance with the Creative Commons Attribution-NonCommercial-NoDerivs 4.0 International License (CC BY-NC-ND 4.0), which permits the non-commercial replication and distribution of the article with the strict proviso that no changes or edits are made and the original work is properly cited (including links to both the formal publication through the relevant DOI and the license). See: https://creativecommons.org/licenses/by-nc-nd/4.0/.
References
- Howard JP, Fisher L, Shun-Shin MJ, et al. Cardiac Rhythm Device Identification Using Neural Networks. JACC Clin Electrophysiol 2019;5:576-86. [Crossref] [PubMed]
- Esteva A, Kuprel B, Novoa RA. Dermatologist-level classification of skin cancer with deep neural networks. Nature 2017;542:115-8. [Crossref] [PubMed]
- Rajpurkar P, Irvin J, Zhu K, et al. CheXNet: Radiologist-Level Pneumonia Detection on Chest X-Rays with Deep Learning. arXiv:1711.05225v3 [cs.CV] 25 Dec 2017.
- Madani A1. Arnaout R2, Mofrad M1, Fast and accurate view classification of echocardiograms using deep learning. NPJ Digit Med 2018; [Crossref]
- Zhang J, Gajjala S, Agrawal PFully Automated Echocardiogram Interpretation in Clinical Practice. Circulation 2018;138:1623-35. [Crossref] [PubMed]
- Zhao Q. py-denseflow. GitHub. Available online: https://github.com/qijiezhao/py-denseflow/
- Donahue J, Hendricks LA, Rohrbach M, et al. Long-Term Recurrent Convolutional Networks for Visual Recognition and Description. IEEE Trans Pattern Anal Mach Intell 2017;39:677-91. [Crossref] [PubMed]
- Tran D, Bourdev L, Fergus R, et al. Learning Spatiotemporal Features with 3D Convolutional Networks. [cited 2019 Apr 24]. Available online: https://arxiv.org/pdf/1412.0767.pdf
- Lin M, Chen Q, Yan S. Network In Network. [cited 2019 Apr 24]. Available online: https://arxiv.org/pdf/1312.4400.pdf
- Carreira J, Zisserman A, Com Z, et al. Quo Vadis, Action Recognition? A New Model and the Kinetics Dataset. [cited 2019 Apr 21]. Available online: https://arxiv.org/pdf/1705.07750.pdf
- Deepmind. I3D models trained on Kinetics. GitHub. 2017. Available online: https://github.com/deepmind/kinetics-i3d
- Feichtenhofer C, Pinz A, Zisserman A. Convolutional Two-Stream Network Fusion for Video Action Recognition. 2016 [cited 2019 Apr 21]. Available online: http://arxiv.org/abs/1604.06573
- Szegedy C, Vanhoucke V, Ioffe S, et al. Rethinking the Inception Architecture for Computer Vision. [cited 2018 Jun 27]. Available online: https://arxiv.org/pdf/1512.00567.pdf
- Abadi M, Agarwal A, Barham P, et al. TensorFlow: Large-Scale Machine Learning on Heterogeneous Distributed Systems. 2016 [cited 2017 Jul 9]. Available online: http://arxiv.org/abs/1603.04467
- Chollet F. Keras: The Python Deep Learning library. 2015; Available online: www.keras.io
- Kotikalapudi R. Keras-vis - Keras Visualization Toolkit. 2018 [cited 2018 Jun 28]. Available online: https://raghakot.github.io/keras-vis/
Cite this article as: Howard JP, Tan J, Shun-Shin MJ, Mahdi D, Nowbar AN, Arnold AD, Ahmad Y, McCartney P, Zolgharni M, Linton NWF, Sutaria N, Rana B, Mayet J, Rueckert D, Cole GD, Francis DP. Improving ultrasound video classification: an evaluation of novel deep learning methods in echocardiography. J Med Artif Intell 2020;3:4.


