
Improving radiograph analysis throughput through transfer learning and object detection
Introduction
Untreated bone fractures can lead to lifelong pain and disability, and treating fractures with traction or plaster casts forces the patient to be immobilized for three months. In contrast, orthopedic surgery enables people with severely broken bones to walk within one week of surgery. For people in low-income countries, this improved time to mobility can prevent families from falling into poverty. For example, in Ghana, 42% of rural households and 40% of urban households experienced a decline in income after the injury of a family member, with 28% of rural households and 19% of urban households reporting an associated decline in food consumption (1). Unfortunately, injury rates are predicted to continue to rise with the increasing use of motorized vehicles, the dominate cause of injury in low- and middle-income countries (LMICs) (2-4).
In comparison to infectious disease, injury treatment is often overlooked by the global health community (5). Surgery is now recognized as a cost-effective intervention for the treatment of injuries (6-8). However, a 2015 Global Health Survey by the Lancet Commission on Global Surgery found that 9 out of 10 people in LMICs do not have access to basic surgical care when needed (7). Thus, there is an urgent need to strengthen the delivery of surgical interventions to treat injuries.
SIGN Fracture Care International is a non-profit organization based in Richland, WA, USA, that helps bridge this gap by working with surgeons in LMICs to administer orthopedic surgery. SIGN provides educational materials and specially designed surgical implants to improve patient outcomes. SIGN donates these implants to partnering hospitals, which then provide treatment at no cost to the patient (9). SIGN’s intramedullary nailing system, designed to treat femoral fractures, has been shown to provide improved healing at reduced cost in limited-resource settings, such as those that lack real-time imaging or power reaming and/or involve delayed presentation to the operating room (10,11).
SIGN-partnered surgeons upload medical data, with imaging typically in the form of radiographs, to allow US-based physicians to provide support to specific cases and evaluate healing. The medical data and corresponding images are organized in SIGN’s Online Surgical Database (SOSD), one of the largest databases on trauma surgery in LMICs (12,13). The SOSD contains over 125,000 cases with 500,000 associated images. This overwhelming amount of medical data presents a challenge to the SIGN team, but also provides an opportunity to make data-driven discoveries that could improve patient care. Pacific Northwest National Laboratory has partnered with SIGN to create computational tools to improve the analytical throughput of radiographs and improve the accuracy of data entered in the SOSD by SIGN-partnered surgeons using computer vision techniques.
Computer vision has dramatically improved over the past several years due to advances in deep learning architectures. In particular, convolutional neural networks (CNNs) can learn hierarchical representations directly from images without relying on handcrafted features, where the deeper the CNN, the greater the level of abstraction of the resulting learned features. Deep CNNs came into vogue after Krizhevsky’s AlexNet, a deep CNN model, greatly outperformed then state-of-the-art computer vision models in the 2010 ImageNet Large-Scale Visual Recognition Challenge (LSVRC) (14). The algorithmic insight inherent in AlexNet was enhanced by the ability to use GPUs to train on a large dataset consisting of 1.2 million images spanning 1,000 categories. This work spawned a variety of deep-CNN-based computer vision models, each competing in various contests to achieve the lowest error rate (15-18).
Girshick et al. showed that a deep CNN architecture can be reformulated for object detection tasks in the 2013 PASCAL Visual Object Classes (VOC) Challenge (19). Originally, the sliding window algorithm was applied for object detection; however, this method was found to be computationally expensive, as the process had to be applied to an image multiple times to detect multiple classes. In current implementations, object detection is achieved by the localization of features in the image using the “recognition using regions” paradigm, which groups pixels into regions via a segmentation algorithm. This method allows encoding of shape and scale information and is not greatly affected by other objects in the background (20). Further improvements in speed and accuracy have been made in recent years (21-23). The TensorFlow team at Google Research has developed an open-source object detection framework built on top of TensorFlow to easily build, train, and deploy such object detection models (24). Identification consists of a bounding box, a label, and a detection score. Figure 1 shows an example of this output for our implant detection model described below.
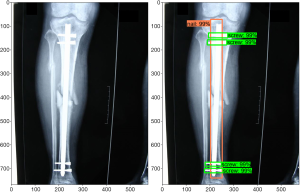
The application of computer vision to medical imaging has generated much interest over the past five years (25,26). Ronneberger et al. developed a deep CNN architecture specifically for biomedical image segmentation called U-Net, which was shown to be applicable to both electron microscopy stacks and transmitted light microscopy images and won the ISBI Cell Tracking Challenge in 2015 (27). Several works have used deep CNNs to segment magnetic resonance images (MRIs) of brains into different tissue classes (28,29). Milletari et al. expanded the use of deep CNNs to the segmentation of MRI volumes of prostates using their V-Net architecture (30). Yu et al. made further advancements in image segmentation for non-medical infrared images through application of a growth immune field, which is described as a combination of immunology and image processing (31).
Though MRI, microscopy, and computed tomography (CT) represent the dominant imaging modalities applied to deep learning thus far, much work has been done concerning radiographs (25). Deep CNNs have been applied to chest radiographs to detect pulmonary tuberculosis and other abnormalities (32-36), knee radiographs to quantify the severity of knee osteoarthritis (37,38), hand radiographs to assess skeletal maturity in children and rheumatoid arthritis in adults (39-41), and wrist and hip radiographs to detect fractures (42-44), among other applications (45-48). The automatic detection of surgical implants in radiographs has not been as thoroughly researched.
Our goal is to automatically detect the number and location of surgical implants in radiographs present in the SOSD. Along with radiographs, images uploaded to the SOSD include surgery photographs and follow-up patient photographs showing clinical function. In order to work only with radiographs, we first developed a machine learning algorithm to sort radiographs—both digital radiographs and photographs of film radiographs—from other types of images.
We then employed the TensorFlow Object Detection API to detect the number and location of surgical implants in the radiographs. Transfer learning using ResNet-50 as the base model was employed to train an implant detection model on a dataset of 2,510 post-op radiographs labeled by bounding boxes. The model trained on the three types of implants under examination (screws, nails, and plates) did a poor job of detecting plates, most likely due to the low proportion of plates in the training set and their varying morphology. Standard image augmentation methods to increase the instance of plates in the training set did not increase the ability of the model to detect plates. Part of this difficulty resulted from the correlation between the plate and screw classes, i.e., plates are held in place by multiple screws, leaning an increase in images with plates to disproportionately increase the instances of screws, disallowing balancing of the classes. We accounted for correlations between plate and screw classes by redrawing the bounding box to include the screws attached to the plate and training two separate models: one to detect nails and screws and one to detect plates. This strategy greatly increased both the precision and recall of plate detection and improved the average precision (AP) by 78.8 percentage points to 92.6%, while the AP for screws and nails remained high at 80.7% and 93.6%, respectively; meanwhile, the sensitivity (true positive rate) was 92% for screws, 86% for nails, and 81% for plates.
Methods
Our work consists of two tasks: classifying images as radiographs and detecting the number and location of surgical implants (screws, nails, and plates) in radiographs. We apply deep learning techniques to accomplish both of these tasks. A major challenge in working with hospitals worldwide is that the radiographs uploaded to the SOSD greatly vary in quality. Digital radiographs are in the minority, and the majority of images are photographs of film radiographs. There are numerous instances of flashes obscuring portions of the radiograph, blurry photos, and visible background, both to the side and through the lighter portions of the radiograph. Because the models developed here are meant to be used in the context of this data, we trained and tested on images of varying quality, removing images from the training and test sets only when a human couldn’t provide a label due to the poor quality of the image.
To sort radiographs from other images, we developed a binary classification model. The image features found by the pre-trained VGG-16 model (16) were extracted, and simple logistic regression was used to perform the binary classification. Logistic regression was implemented using Scikit-learn (49) with L2 regularization and the LIBLINEAR solver (50). Because radiographs tend to have high contrast and are mostly black and white, while non-radiographic images in the SOSD tend to be full color and have less stark contrast, we applied image manipulation strategies to better balance the color space. We duplicated the non-radiographic images in our training set and inverted the colors, then duplicated that collection and made the images monochrome to ensure our model was not simply learning color gradients. This produced 5,151 radiographs and 3,994 non-radiographic images for use as the training set. From this, 10% of the images were set aside as a validation set to check the accuracy of our model. A test set containing 683 radiographs and 111 non-radiographic images was created to examine the classification ability of our model on unaltered images in the SOSD.
For the detection of surgical implants, we applied the TensorFlow Object Detection API. In the TensorFlow Object Detection API, pre-trained models from the TensorFlow Model Zoo are applied to the detection of objects in widely used image datasets, such as the COCO dataset (51). This image dataset, like other prevalent image datasets, consists of commonly identifiable objects in images found on the Internet. Naturally, this excludes radiographs and surgical implants. Therefore, we applied transfer learning to leverage pre-optimized computer vision models for the development of a model that detects surgical implants in radiographs. Such domain-specific fine-tuning has been proven to be an effective strategy for training high-capacity CNNs when data is scarce (19), with Tajbakhsh et al. showing that fine-tuning is also acceptable for a variety of medical imaging data (52). Though the SOSD contains a wealth of radiographs, labeling each image with bounding boxes surrounding implants in the three classes under study (nails, screws, and plates) is time-consuming; therefore, our implant detection model can benefit from the utilization of fine-tuning techniques.
In our implant detection model, the training dataset was generated by overlaying boxes on each implant in the radiographs. These boxes act as anchor points on which the model is trained. We used the open-source graphical image annotation tool LabelImg to create XML files that contained information about the box location and label of implants in the sample radiographs. These files were then converted into a single TensorFlow record file to be read into the Object Detection API.
One can imagine that hand-labeling thousands of radiographs would be quite tedious. To speed the labeling process, we applied an active learning technique. An initial model was fine-tuned on 370 hand-labeled radiographs leveraging the weights from Faster R-CNN ResNet-50 trained on the COCO dataset. This model was then used to detect implants in a new set of radiographs from the SOSD, with a detection threshold of 88%. All detected objects were checked for fidelity, and any anchoring errors or missing identifications were corrected. These new images were incorporated in the training set, and the model was retrained. Repeating this cycle three times allowed us to more quickly generate a training dataset containing 2,510 labeled images.
The object detection model was validated by computing the average precision (AP) and true and false positive rates of each class (53). Determining the AP of a class in an object detection model relies on the intersection over union (IoU) to label the detections as true or false positives. Typically, a detection is labeled a true positive if the IoU is greater than or equal to 0.5. Using this value, we discovered that our model suffered from decreased localization. This led to true positive detections being reported as false positives when an extended portion of the implant was left out of the bounding box. To overcome this issue, we decreased the IoU threshold for true positive detections to 0.35. For the final version of the model (discussed below), the mean average precision (mAP) was 82.4% with an IoU of 0.5, while decreasing the IoU to 0.35 improved the mAP to 92.5%, and the false positive rate decreased from 15.9% to 14.0%. Therefore, when computing the AP and true and false positive rates, we used an IoU of 0.35.
Results and discussion
To work with radiographs in the SOSD, we first developed a classification model to sort radiographs from non-radiographic images. We applied transfer learning using the VGG-16 model, training on 8,230 images and validating with 915 images. After feature extraction and logistic regression, 100% of the validation images were correctly classified. From a test set of 683 radiographs and 111 non-radiographic images, only 1 image was misclassified: a photograph of a leg was classified as a radiograph. The receiver operating characteristic (ROC) curve on the test set is shown in Figure 2. The dotted line represents classification by pure chance, while the solid blue line represents classification by our model. Our model shows near-perfect classification, with an area under the curve (AUC) of 0.995.
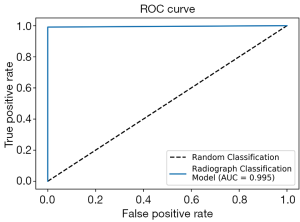
After removing non-radiographic images from our image set, we developed an object detection model to detect surgical implants in radiographs. The radiographs in the SOSD contain three types of implants: nails, screws, and plates. Our initial object detection model was trained to detect all three implants simultaneously. Typically, when training object detection models, all classes should have approximately the same number of instances in the training set. However, because we do not have isolated instances of each class, it becomes more difficult to curate a training set with comparable instances of each class. Each nail is secured with a variable number of screws (typically 1–4), and each plate is secured with a large number of screws (5+). Thus, every time we add radiographs with nails or plates to the training set, we inevitably disproportionately increase the instances of screws. In addition, because SIGN designs and distributes nails to SIGN-partnered surgeons, the SOSD contains many more radiographs containing nails than plates. Our initial training set of 2,510 images contained 10,558 screws, 3,734 nails, and 797 plates. Such a skewed dataset is typically undesirable in deep learning methods, and often the training set is balanced by applying image manipulations to classes with low representation. However, as stated, the three classes in this dataset are not independent of one another. Of the 2,446 images containing nails, 99% also contain screws. Meanwhile, 100% of the 564 images that contain plates also contain screws, while 92% of the images with plates also contain nails. Therefore, we cannot simply apply image augmentation techniques to increase the count of objects in one class without increasing the count of objects in the other classes.
To confirm that image augmentation techniques are not applicable in the case of associated classes, we manipulated the radiographs containing plates to double the representation of plates in the training set. Though it is not possible to completely balance the dataset so that each class is represented equally, we were able to increase the instance of plates relative to nails. To 292 radiographs containing plates, we applied either a horizontal flip, vertical flip, 90° rotation, 180° rotation, or 270° rotation; the aspect ratio of some of the rotated radiographs were also manipulated to add further distortions. In addition, we applied color inversions and/or 90° rotations to 508 images from the full training set. After these manipulations, our augmented dataset was composed of 5,443 images, containing 30,622 screws (approx. 5.6 per image), 8,475 nails (approx. 1.5 per image), and 4,226 plates (approx. 0.8 per image).
Precision–recall curves for each implant class are shown in Figure 3, with the average precision (AP) given in the legend. Precision is defined as the ratio of true positives to all predicted positives, and recall is defined as the ratio of true positives to the total number of ground truth objects in that class. Ideally, a model would have both high precision and high recall; however, in practice, there is typically a tradeoff between precision and recall. This trade-off is present in our case for all three classes, as shown by the decrease in precision as recall increases.
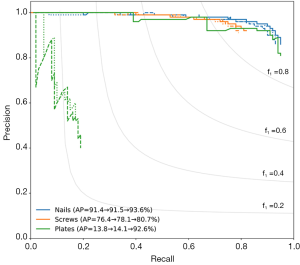
When training the model without image augmentation (dotted lines in Figure 3), the AP was 91.4% for nails, 76.4% for screws, and 13.8% for plates. While this model did well at detecting nails and screws, plate scores were inadequate, likely due to the low instance of plates in the training set, as discussed above. The model with image augmentation applied (dashed lines in Figure 3) showed insignificant improvement, with an AP of 91.5% for nails, 78.1% for screws, and 14.1% for plates.
Plate detection is likely a more difficult task than screw or nail detection simply because of the variety of orthopedic plates used in practice. The type of fixation device is highly dependent on the bone under repair and the type of fracture. To accommodate the specific need, orthopedic plates come in a variety of shapes and with variable numbers of holes. Many tend to be symmetrical with variable width, while other others, such as buttress plates, are often asymmetrical. Depending on the bone, plates can be curved or straight, and sometimes multiple plates are used in conjunction. For surgical screws, though the length, diameter, threading, and bolt-head dimensions may vary, the overall appearance is consistent. In this dataset, the majority of nails were provided to the surgeons by SIGN and tend to differ mainly in length and shape of the ends, but again the overall appearance is consistent. Therefore, we trained a separate model to detect only plates. The bounding boxes for plates were redrawn to include the screws used to hold the plates in place on 547 radiographs from the SOSD (see Figure 4 for comparison of the bounding boxes). Five hundred of these images were used to train the plate-only detection model, while the remaining 47 images were used to test the model. A separate model was trained on the images with bounding boxes described previously to detect only screws and nails (2,510 images total). To test this model, a set of 1,000 images was obtained from the SOSD. This set was labeled using the active learning method described above to generate ground truth bounding boxes. The test set contained 930 nails and 2,165 screws.
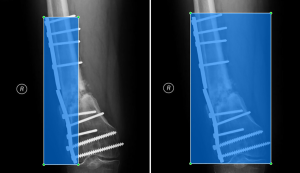
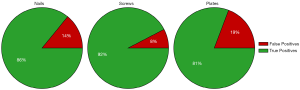
Training two separate models (one to detect nails and screws, and one to detect plates) greatly improved both the precision and recall of plate detection, as evidenced by the solid green lines in Figure 3. The AP of the plate dramatically increased to 92.6% when a separate model was used. Separating the models also had a modest positive effect on nail and screw detection. The class AP increased from 91.4% to 93.6% for nails and from 76.4% to 80.7% for screws. In addition, Figure 5 shows true and false detection rates for each class from the model trained on screws and nails and the model trained on plates. The true positive rate, also called the sensitivity, is highest for screws at 92%, followed by nails at 86%, and plates at 81%. This indicates that although the precision was lower for screws, detections in this class are more likely to be labeled correctly.
Therefore, producing a separate model for the detection of low-representation objects is a good strategy for improving object detection when objects are highly correlated in the images and standard image augmentation techniques do not improve detection. Furthermore, considering these correlations when labeling objects (i.e., a plate will always be accompanied by screws) also improves the model by changing the representation to include correlated objects.
Conclusions
SIGN works with 52 hospitals around the world to serve thousands of patients. This work has allowed SIGN to build a database consisting of over 500,000 images, which, if analyzed, could generate data-driven conclusions on how to improve patient outcomes. However, because SIGN-partnered surgeons are located in different regions and have different available resources, the medical images uploaded to the SOSD are of varying quality. Most radiographs provided to the SOSD are photographs of film radiographs, and there is no standard method or equipment used to take the photographs. This large variety makes the use of standard analysis techniques difficult. To overcome this challenge, we apply deep learning techniques to the analysis of radiographs.
There is a wealth of tools available for automatic image analysis, a number of which are based in deep learning. Many of these tools are available on public repositories and, with some effort, can be tuned for highly specific applications, in this case, the identification of surgical implants in radiographs. Generalized models trained on millions of images to detect hundreds of classes of objects can be fine-tuned for a specific use with a relatively low number of images. In this work, we fine-tuned the Faster R-CNN ResNet-50 model pretrained on the ImageNet dataset to detect surgical nails, screws, and plates in radiographs from the SOSD. Because surgical implants are often used in conjunction, it was not possible to create a balanced dataset, and attempting to improve the balance of plates through standard image augmentation techniques did not improve the precision or recall of the plate class. We improved plate detection by considering correlations between the plate and screw classes. We redrew the bounding boxes around the plates to include the screws used to hold the plate in place and used these images to train a model that detects plates. This strategy increased the AP of plate detection by 78.8 percentage points. In sum, the AP of each class was 80.7% for screws, 93.6% for nails, and 92.6% for plates, while the sensitivity was 92% for screws, 86% for nails, and 81% for plates.
Finally, we ran our object detection model over all radiographs in the SOSD to correct erroneous entries. The results from the object detection tool will be used in future work aiming to suggest optimal surgical parameters based on the type and location of femoral fracture using information on patient outcomes from the SOSD. We are currently working with SIGN to implement these tools on their servers to allow radiographs uploaded in the future to be quickly assessed for type, location, and number of hardware present.
Acknowledgments
The authors thank Tonya Martin, Carlos Ortiz Marrero, Michael Girard, Tom Warfel, and Robert Rallo for helpful discussions.
Funding: The research described in this paper was conducted under the Laboratory Directed Research and Development Program at Pacific Northwest National Laboratory, a multiprogram national laboratory operated by Battelle for the U.S. Department of Energy (Release No. PNNL-SA-149813).
Footnote
Conflicts of Interest: All authors have completed the ICMJE uniform disclosure form (available at http://dx.doi.org/10.21037/jmai-20-2). The authors have no conflicts of interest to declare. SIGN Fracture Care International is registered as a nonprofit, tax-exempt corporation in the state of Washington and in the United States with IRS 501(c)(3) status.
Ethical Statement: The authors are accountable for all aspects of the work in ensuring that questions related to the accuracy or integrity of any part of the work are appropriately investigated and resolved.
Open Access Statement: This is an Open Access article distributed in accordance with the Creative Commons Attribution-NonCommercial-NoDerivs 4.0 International License (CC BY-NC-ND 4.0), which permits the non-commercial replication and distribution of the article with the strict proviso that no changes or edits are made and the original work is properly cited (including links to both the formal publication through the relevant DOI and the license). See: https://creativecommons.org/licenses/by-nc-nd/4.0/.
References
- Mock CN, Gloyd S, Adjei S, et al. Economic consequences of injury and resulting family coping strategies in Ghana. Accid Anal Prev 2003;35:81-90. [Crossref] [PubMed]
- Murray CJL, Vos T, Lozano R, et al. Disability-adjusted life years (DALYs) for 291 diseases and injuries in 21 regions, 1990-2010: a systematic analysis for the global burden of disease study 2010. Lancet 2012;380:2197-223. [Crossref] [PubMed]
- WHO. Global status report on road safety: time for action World Health Organization. Geneva: World Health Organization, 2009.
- Patton GC, Coffey C, Sawyer SM, et al. Global patterns of mortality in young people: a systematic analysis of population health data. Lancet 2009;374:881-92. [Crossref] [PubMed]
- Gosselin RA, Spiegel DA, Coughlin R, et al. Injuries: the neglected burden in developing countries. Bull World Health Organ 2009;87:246-246a. [Crossref] [PubMed]
- Gosselin RA, Heitto M. Cost-effectiveness of a district trauma hospital in Battambang, Cambodia. World J Surg 2008;32:2450. [Crossref] [PubMed]
- Meara JG, Leather AJM, Hagander Lo. Global surgery 2030: Evidence and solutions for achieving health, welfare, and economic development. Lancet 2015;386:569-624. [Crossref] [PubMed]
- Mock CN, Donkor P, Gawande A, et al. Essential surgery: Key messages from disease control priorities. Lancet 2015;385:2209-19. [Crossref] [PubMed]
- Zirkle LG. Injuries in developing countries—how can we help? The role of orthopaedic surgeons. Clin Orthop Relat Res 2008;466:2443. [Crossref] [PubMed]
- Phillips J, Zirkle LG, Gosselin RA. Achieving locked intramedullary fixation of long bone fractures: Technology for the developing world. Int Orthop 2012;36:2007-13. [Crossref] [PubMed]
- Sekimpi P, Okike K, Zirkle L, et al. Femoral fracture fixation in developing countries: An evaluation of the surgical implant generation network (SIGN) intramedullary nail. J Bone Joint Surg Am 2011;93:1811-8. [Crossref] [PubMed]
- Clough JF, Zirkle LG, Schmitt RJ. The role of sign in the development of a global orthopaedic trauma database. Clin Orthop Relat Res 2010;468:2592-7. [Crossref] [PubMed]
- Young S, Lie SA, Hallan Go. Risk factors for infection after 46,113 intramedullary nail operations in low- and middle-income countries. World J Surg 2013;37:349-55. [Crossref] [PubMed]
- Krizhevsky A, Sutskever I, Hinton GE. ImageNet classification with deep convolutional neural networks. Communications of the ACM 2017;60:84-90. [Crossref]
- He K, Zhang X, Ren S, et al. Deep residual learning for image recognition. In 2016 IEEE Conference on Computer Vision and Pattern Recognition (CVPR), 2016:770-8.
- Simonyan K, Zisserman A. Very Deep Convolutional Networks for Large-Scale Image Recognition. In Proceedings of the 3rd International Conference on Learning Representations, 2015.
- Szegedy C, Wei Liu, Yangqing Jia, et al. Going deeper with convolutions. In 2015 IEEE Conference on Computer Vision and Pattern Recognition (CVPR), 2015:1-9.
- Szegedy C, Vanhoucke V, Ioffe S, et al. Rethinking the Inception Architecture for Computer Vision. In Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition, 2016:2818-26.
- Girshick RB, Donahue J, Darrell T, et al. Rich feature hierarchies for accurate object detection and semantic segmentation. In Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition, 2014:580-7.
- Chunhui Gu, Lim JJ, Arbelaez P, et al. Recognition using regions. In 2009 IEEE Conference on Computer Vision and Pattern Recognition, 2009:1030-7.
- Liu W, Anguelov D, Erhan D, et al. SSD: single shot multibox detector In Computer Vision-ECCV 2016:21-37.
- Redmon J, Divvala S, Girshick R, et al. You only look once: Unified, real-time object detection In Proceedings of the IEEE conference on computer vision and pattern recognition, 2016:779-88.
- Ren S, He K, Girshick R, et al. Faster r-CNN: Towards real-time object detection with region proposal networks. In Advances in neural information processing systems, 2015:91-9.
- Huang J, Rathod V, Sun C, et al. Speed/accuracy trade-offs for modern convolutional object detectors In 2017 IEEE Conference on Computer Vision and Pattern Recognition (CVPR), 2017:3296-7.
- Litjens G, Kooi T, Bejnordi BE, et al. A survey on deep learning in medical image analysis. Medical Image Analysis 2017;42:60-88. [Crossref] [PubMed]
- Shen D, Wu G, Suk HI. Deep learning in medical image analysis. Annu Rev Biomed Eng 2017;19:221-48. [Crossref] [PubMed]
- Ronneberger O, Fischer P, Brox T. U-net: Convolutional networks for biomedical image segmentation. In: Navab N, Hornegger J, Wells WM, et al. editors. Medical Image Computing and Computer-Assisted Intervention-MICCAI. Cham. Springer International Publishing, 2015:234-41.
- Moeskops P, Viergever MA, Mendrik AM, et al. Automatic segmentation of MR brain images with a convolutional neural network. IEEE Trans Med Imaging 2016;35:1252-61. [Crossref] [PubMed]
- Zhang W, Li R, Deng H, et al. Deep convolutional neural networks for multi-modality isointense infant brain image segmentation. Neuroimage 2015;108:214-24. [Crossref] [PubMed]
- Milletari F, Navab N, Ahmadi S. V-net: Fully convolutional neural networks for volumetric medical image segmentation In 2016 Fourth International Conference on 3D Vision (3DV), 2016:565-71.
- Yu X, Zhou Z, Gao Q, et al. Infrared image segmentation using growing immune field and clone threshold. Infrared Physics & Technology 2018;88:184-93. [Crossref]
- Annarumma M, Withey SJ, Bakewell RJ, et al. Automated triaging of adult chest radiographs with deep artificial neural networks. Radiology 2019;291:196-202. [Crossref] [PubMed]
- Cicero M, Bilbily A, Colak E, et al. Training and validating a deep convolutional neural network for computer-aided detection and classification of abnormalities on frontal chest radiographs. Invest Radiol 2017;52:281-7. [Crossref] [PubMed]
- Lakhani P, Sundaram B. Deep learning at chest radiography: Automated classification of pulmonary tuberculosis by using convolutional neural networks. Radiology 2017;284:574-82. [Crossref] [PubMed]
- Rajkomar A, Lingam S, Taylor AG, et al. High-throughput classification of radiographs using deep convolutional neural networks. J Digit Imaging 2017;30:95-101. [Crossref] [PubMed]
- Yang W, Chen Y, Liu Y, et al. Cascade of multi-scale convolutional neural networks for bone suppression of chest radiographs in gradient domain. Med Image Anal 2017;35:421-33. [Crossref] [PubMed]
- Antony J, McGuinness K, O’Connor NE, et al. Quantifying radiographic knee osteoarthritis severity using deep convolutional neural networks In 2016 23rd International Conference on Pattern Recognition (ICPR), 2016:1195-200.
- Norman B, Pedoia V, Noworolski A, et al. Applying densely connected convolutional neural networks for staging osteoarthritis severity from plain radiographs. J Digit Imaging 2019;32:471-7. [Crossref] [PubMed]
- Iglovikov VI, Rakhlin A, Kalinin AA, et al. (2018) Paediatric bone age assessment using deep convolutional neural networks In Stoyanov D, Taylor Z, Carneiro G, et al. editors. Deep Learning in Medical Image Analysis and Multimodal Learning for Clinical Decision Support. Cham. Springer International Publishing, 2018:300-8.
- Larson DB, Chen MC, Lungren MP, et al. Performance of a deep-learning neural network model in assessing skeletal maturity on pediatric hand radiographs. Radiology 2018;287:313-22. [Crossref] [PubMed]
- Üreten K, Erbay H, Maras HH. Detection of rheumatoid arthritis from hand radiographs using a convolutional neural network. Clin Rheumatol 2020;39:969-74. [Crossref] [PubMed]
- Gan K, Xu D, Lin Y, et al. Artificial intelligence detection of distal radius fractures: a comparison between the convolutional neural network and professional assessments. Acta Orthop 2019;90:394-400. [Crossref] [PubMed]
- Kim DH, MacKinnon T. Artificial intelligence in fracture detection: transfer learning from deep convolutional neural networks. Clin Radiol 2018;73:439-45. [Crossref] [PubMed]
- Urakawa T, Tanaka Y, Goto S, et al. Detecting intertrochanteric hip fractures with orthopedist-level accuracy using a deep convolutional neural network. Skeletal Radiol 2019;48:239-44. [Crossref] [PubMed]
- Aubert B, Vidal PA, Parent S, et al. Convolutional neural network and in-painting techniques for the automatic assessment of scoliotic spine surgery from biplanar radiographs In: Descoteaux M, Maier-Hein L, Franz A, et al. editors, Medical Image Computing and Computer-Assisted Intervention-MICCAI. Cham. Springer International Publishing, 2017:691-9.
- Cheng PM, Tejura TK, Tran KN, et al. Detection of high-grade small bowel obstruction on conventional radiography with convolutional neural networks. Abdom Radiol (NY) 2018;43:1120-7. [Crossref] [PubMed]
- Pelka O, Nensa F, Friedrich CM. Annotation of enhanced radiographs for medical image retrieval with deep convolutional neural networks. PLoS One 2018;13:e0206229. [Crossref] [PubMed]
- Togo R, Yamamichi N, Mabe K, et al. Detection of gastritis by a deep convolutional neural network from double-contrast upper gastrointestinal barium X-ray radiography. J Gastroenterol 2019;54:321-9. [Crossref] [PubMed]
- Pedregosa F, Varoquaux G, Gramfort A, et al. Scikit-learn: Machine learning in Python. J Chem Inf Model 2011;12:2825-30.
- Fan RE, Chang KW, Hsieh CJ, et al. LIBLINEAR: A library for large linear classification. J Chem Inf Model 2008;9:1871-4.
- Lin TY, Maire M, Belongie S, et al. Microsoft coco: Common objects in context In Computer Vision-ECCV. Cham. Springer International Publishing, 2014:740-55.
- Tajbakhsh N, Shin JY, Gurudu SR, et al. Convolutional neural networks for medical image analysis: Full training or fine tuning? IEEE Trans Med Imaging 2016;35:1299-312. [Crossref] [PubMed]
- Cartucho J, Ventura R, Veloso M. Robust object recognition through symbiotic deep learning in mobile robots In 2018 IEEE/RSJ International Conference on Intelligent Robots and Systems (IROS), 2018:2336-41.
Cite this article as: Bilbrey JA, Ramirez EF, Brandi-Lozano J, Sivaraman C, Short J, Lewis ID, Barnes BD, Zirkle LG. Improving radiograph analysis throughput through transfer learning and object detection. J Med Artif Intell 2020;3:9.


